- Blog
- Sony vegas movie studio hd platinum 11 torrenty-org
- Adobe media encoder software performance encoding baseline
- Best free 3d racing games download for pc
- Adobe photoshop cc reviews
- Microsoft office academic calendar templates
- Cs go bunnyhop script download
- Ezdrummer 2 keygen patch
- Insert current date in excel document
- Dear zindagi movie in full hd download
- Pretty borders for word documents
- Adobe photoshop cs6 torrent file
- Invoice for mac free word
- Install soundflower mac os x lion 10-7-5
- Latin modern roman for word
- Does r strudio come with r markdown for mac

- #Does r strudio come with r markdown for mac how to#
- #Does r strudio come with r markdown for mac code#
- #Does r strudio come with r markdown for mac download#
A popup will show you what the resulting document looks like. This will execute the R Markdown file and create the file “test.html” in your folder. When you’re done editing, click the “Knit” at the top of the window with your R Markdown script.
#Does r strudio come with r markdown for mac code#
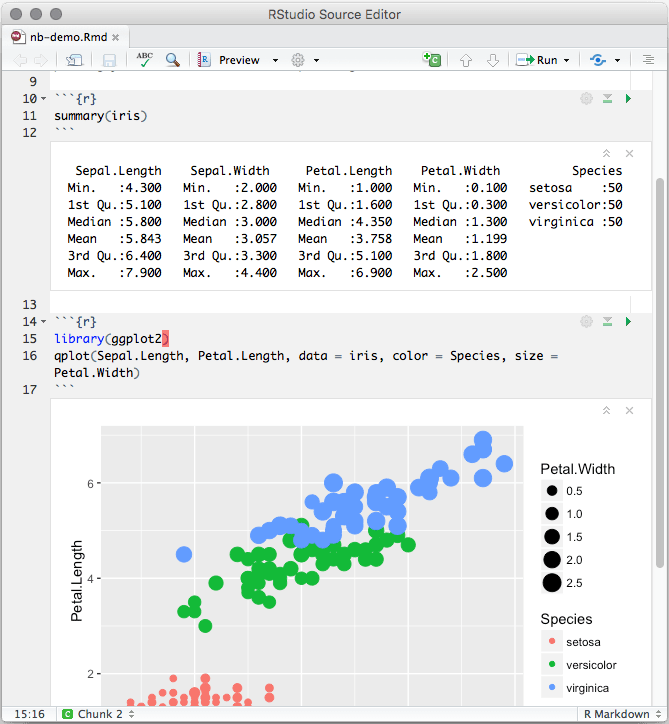
This can be helpful if you want only to make a point or to show a variation of R code that also works but you don’t want to clutter the resulting html file with the output. You don’t need to specify include = TRUE, to include the output of the code in the results, because it is the default.Įvery once in while you might want to use eval = FALSE in a code chunk instead, to prevent the code chunk in your document from being executed. Put eval = TRUE there so that code in this chunk will be executed. This code chunk also includes options between curly brackets. You can modify this as you learn more about this and other global options in RMarkdown. This first code chunk will contain global instructions – here, the only global instruction is to set the size of the figures produced in the output file. Code chunks begin and end between triple single backquotes. The next part of your R Markdown file is an R code chunk. These instructions are to save your output to an html file and to include a floating table of contents. For now, change only the title between the quotes and leave the rest of the code as-is. The section begins and ends with lines containing three dashes ‘-’ and sets output parameters (file type, document title, author name, table of contents style, etc.). The top of the opened R Markdown file should look like the following. This will open a R Markdown script file for editing in R Studio.
Rmd file you renamed it) in the Files pane of your R Studio session. Click on the file “test.Rmd” (or whatever.Proj file to reopen the project in R Studio. From now on you can just double click the. In the Files pane of your R Studio session, you should see the files you saved to the new folder and the “.Rproj” file just created. R will create a “.Proj” project file in your new folder. Choose “Existing Directory” and then navigate to your new folder.Run R Studio and select “File” -> “New Project” from the top menu.The easiest way to continue with R Markdown is to use RStudio. You can duplicate and rename test.Rmd whatever you like (e.g., Assignment_1.Rmd), but keep the other file names as they are.
#Does r strudio come with r markdown for mac download#
Download the following three files and save one at a time to your new folder (right-click and save to disk): test.Rmd, mystyles.css, and _site.yml.Create a new folder on your computer to contain your R Markdown documents.You might find this cheat sheet useful once you are up and running. Visit the R Studio R Markdown help pages for more organized information on R Markdown.
#Does r strudio come with r markdown for mac how to#
Below is a minimal description of how to get started.Īs usual, Google is your source for answers. Try using R Markdown to make documents that combine text and executable R code. The following example changes the trim option to calculate a trimmed mean, mean(myvariable, trim = 0.1) (If you fail to do this and have missing values, R will return NA.) mean(myvariable, na.rm = TRUE) This instruct R to remove missing values from the data object before calculating the mean. The following example changes the na.rm option to TRUE. If the default values for the options don’t meet your needs you can alter the values. If you want the mean of the elements in the variable myvariable, enter mean(myvariable) If you are happy with the default settings, then you can use the command in its simplest form. In the case of the mean almost any data object will do, but you will usually apply the function to a vector (representing a single variable). Look under “Arguments” on the help page to see what kind of object R needs. In this example, the argument x represents the data object you supply to the function. Any argument with an “=” sign represents an option, with the default value indicated. The items between the brackets “()” are called arguments.Īny argument without an “=” sign is required - you must provide it for the command to work. In the pop-up help window, look under the title “Usage” and you will see something like this: mean(x, trim = 0, na.rm = FALSE. As an example, here’s how to interpret the help page for the sample mean, obtained by ?mean
